 Compressed index
Compressed index
Participants:
Chandana, Jing Quan Lim, Hoang, Wing-Kai Hon, Tak-Wah Lam, Sadakane, Wing-Kin Sung
Background
DNA sequences, which hold the code of life for every living organism, can be represented by strings over 4 characters A, C, G, and T.
Due to the advance in bio-technology, we already know the complete sequences for a number of living organisms, including human.
The advance in sequencing also generated many sequencing data.
We faced two problems. First, the dataset is too big. It is easily generate hundreds of billions of data per individual.
Second, to do analysis, biologists require tools that can locate the positions of an arbitrary pattern over a long DNA sequence efficiently.
However, genomic data is long. It is time consuming to search a parttern by linearly scan the DNA sequence.
Objectives
To resolve the issue of hugh amount of data, we can use compression techniques.
To resolve the pattern searching problem, we can use data-structure.
Now, we need to resolve two issues at the same time.
In this project, we aim to create indexing data-structure that is compressed. There ae a few subproblems.
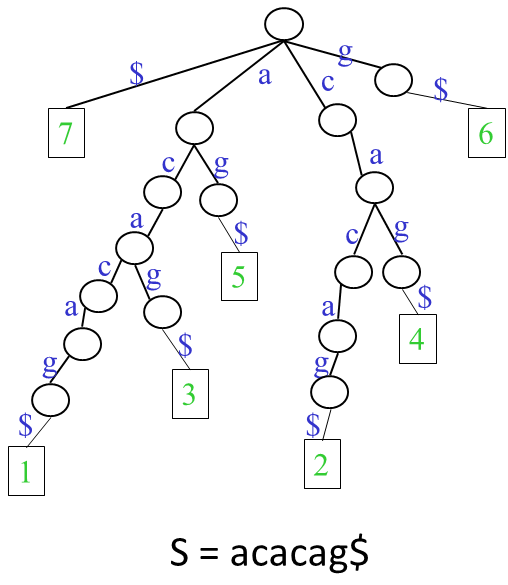
- Constructing compressed suffix array efficiently and space-efficiently: For pattern matching, the most useful data-structures are suffix tree and suffix array. They enable fast pattern matching. The only issue is that they are very big. Due to the works of Grossi, Vitter, Ferragina and Manzini, we can have full-text index whose size is as small as n Hk + o(n) bits. However, when the text is long, it is not possible to construct them space-efficiently. This project is aimed to create some fast algorithms for constructing BWT, FM-index or compressed suffix array. We proposed a method that runs in O(n) time and uses O(n) bits space. Later, based on the success, we gave a practically fast solution. Using our techniques, we can construct the suffix array for the whole human genome in less than 1 hour in a Personal Computer.
- Efficient algorithm for searching patterns allowing errors: Due to sequencing errors and mutations, the reads generated by the sequencing machine do not exactly match the reference human genome. Hence, when we perform pattern matching, we need to allow mismatches/errors. In this project, we developed a number of different techniques for efficient pattern matching.
- Applying compressed data-structure in next generation seqeuncing: Based on the results of the first two sub-projects, we applied these algorithms for next generation sequencing analysis. We developed
two general aligners BatMis (Bioinformatics 2012) and BatAlign (NAR 2015). We also developed a method BatMeth (Genome Biology 2012) for aligning bisulfite reads.
Software
- BatMeth: Improved mapper for bisulfite sequencing reads on DNA methylation
- BatMis: A fast short read aligner allowing up to 10 mismatches
- BatAlign: an incremental method for accurate alignment of sequencing reads
Selected Publications
-
Lim, J.-Q., Tennakoon, C., Guan, P. and Sung, W.-K.
BatAlign: an incremental method for accurate alignment of sequencing reads.
Nucleic Acids Res,
2015.
-
Do, H. H., Jansson, J., Sadakane, K. and Sung, W.
Fast relative Lempel-Ziv self-index for similar sequences.
Theor. Comput. Sci.,
Vol. 532, pp. 14-30, 2014.
-
Do, H. H. and Sung, W.-K.
Compressed Directed Acyclic Word Graph with Application in Local Alignment.
Algorithmica,
Vol. 67(2), pp. 125-141, 2013.
-
Lim, J.-Q., Tennakoon, C., Li, G., Wong, E., Ruan, Y., Wei, C.-L. and Sung, W.-K.
BatMeth: improved mapper for bisulfite sequencing reads on DNA methylation.
Genome Biol,
Vol. 13(10), pp. R82, 2012.
-
Tennakoon, C., Purbojati, R. W. and Sung, W.-K.
BatMis: A fast algorithm for k-mismatch mapping.
Bioinformatics,
2012.
-
Jansson, J., Sadakane, K. and Sung, W.-K.
CRAM: Compressed Random Access Memory.
ICALP (1),
510-521, 2012.
-
Jansson, J., Sadakane, K. and Sung, W.-K.
Ultra-succinct representation of ordered trees with applications.
J. Comput. Syst. Sci.,
Vol. 78(2), pp. 619-631, 2012.
-
Chan, H.-L., Lam, T. W., Sung, W.-K., Tam, S.-L. and Wong, S.-S.
A linear size index for approximate pattern matching.
J. Discrete Algorithms,
Vol. 9(4), pp. 358-364, 2011.
-
Chan, H.-L., Lam, T.-W., Sung, W.-K., Tam, S.-L. and Wong, S.-S.
Compressed Indexes for Approximate String Matching.
Algorithmica,
Vol. 58(2), pp. 263-281, 2010.
-
Hon, W.-K., Sadakane, K. and Sung, W.-K.
Breaking a Time-and-Space Barrier in Constructing Full-Text Indices.
SIAM J. Comput.,
Vol. 38(6), pp. 2162-2178, 2009.
-
Lam, T. W., Sung, W.-K., Tam, S.-L., Wong, C.-K. and Yiu, S.-M.
Compressed indexing and local alignment of DNA.
Bioinformatics,
Vol. 24(6), pp. 791-797, 2008.
-
Lam, T. W., Sung, W.-K. and Wong, S.-S.
Improved Approximate String Matching Using Compressed Suffix Data Structures.
Algorithmica,
Vol. 51(3), pp. 298-314, 2008.
-
Hon, W.-K., Lam, T. W., Sadakane, K., Sung, W.-K. and Yiu, S.-M.
A Space and Time Efficient Algorithm for Constructing Compressed Suffix Arrays.
Algorithmica,
Vol. 48(1), pp. 23-36, 2007.
-
Jansson, J., Sadakane, K. and Sung, W.-K.
Compressed Dynamic Tries with Applications to LZ-Compression in Sublinear Time and Space.
FSTTCS,
Vol. 4855, pp. 424-435, 2007.
-
Huynh, T. N. D., Hon, W.-K., Lam, T. W. and Sung, W.-K.
Approximate string matching using compressed suffix arrays.
Theor. Comput. Sci.,
Vol. 352(1-3), pp. 240-249, 2006.
-
Hon, W.-K., Lam, T. W., Sadakane, K. and Sung, W.-K.
Constructing Compressed Suffix Arrays with Large Alphabets.
ISAAC,
240-249, 2003.
-
Hon, W.-K., Lam, T. W., Sadakane, K., Sung, W.-K. and Yiu, S.-M.
Compressed Index for Dynamic Text.
Data Compression Conference,
102-111, 2004.
-
Hon, W.-K., Lam, T. W., Sung, W.-K., Tse, W.-L., Wong, C.-K. and Yiu, S.-M.
Practical aspects of Compressed Suffix Arrays and FM-Index in Searching DNA Sequences.
ALENEX/ANALC,
31-38, 2004.
-
Huynh, T. N. D., Hon, W.-K., Lam, T. W. and Sung, W.-K.
Approximate String Matching Using Compressed Suffix Arrays.
CPM,
434-444, 2004.
-
Hon, W.-K., Sadakane, K. and Sung, W.-K.
Breaking a Time-and-Space Barrier in Constructing Full-Text Indices.
FOCS,
251-260, 2003.
-
Lam, T. W., Sadakane, K., Sung, W.-K. and Yiu, S.-M.
A Space and Time Efficient Algorithm for Constructing Compressed Suffix Arrays.
COCOON
401-410, 2002.
-
Lam, T. W., Sung, W.-K., Tam, S.-L., Wong, C.-K. and Yiu, S.-M.
An Experimental Study of Compressed Indexing and Local Alignments of DNA.
COCOA
242-254, 2007.
-
Lam, T. W., Sung, W.-K., Tam, S.-L. and Yiu, S.-M.
Space Efficient Indexes for String Matching with Don't Cares.
ISAAC,
846-857, 2007.
Last updated: 30/12/2015, Wing-Kin Sung.
 Compressed index
Compressed index
 Compressed index
Compressed index